Sitemap
A list of all the posts and pages found on the site. For you robots out there, there is an XML version available for digesting as well.
Pages
Posts
Future Blog Post
Published:
This post will show up by default. To disable scheduling of future posts, edit config.yml and set future: false.
Blog Post number 4
Published:
This is a sample blog post. Lorem ipsum I can’t remember the rest of lorem ipsum and don’t have an internet connection right now. Testing testing testing this blog post. Blog posts are cool.
Blog Post number 3
Published:
This is a sample blog post. Lorem ipsum I can’t remember the rest of lorem ipsum and don’t have an internet connection right now. Testing testing testing this blog post. Blog posts are cool.
Blog Post number 2
Published:
This is a sample blog post. Lorem ipsum I can’t remember the rest of lorem ipsum and don’t have an internet connection right now. Testing testing testing this blog post. Blog posts are cool.
Blog Post number 1
Published:
This is a sample blog post. Lorem ipsum I can’t remember the rest of lorem ipsum and don’t have an internet connection right now. Testing testing testing this blog post. Blog posts are cool.
portfolio
Adaptation of genetic evolution methods to studying cultural evolution
Cultural evolution is the study of how socially learned behaviors, ideas, technologies, and norms change over time by processes analogous to biological evolution, but operating through social learning rather than genes. I study cultural variation using methods inspired by biological evolution, including the extraction of hierarchical structure in cultural variation and the intergenerational transmission of cultural traits.
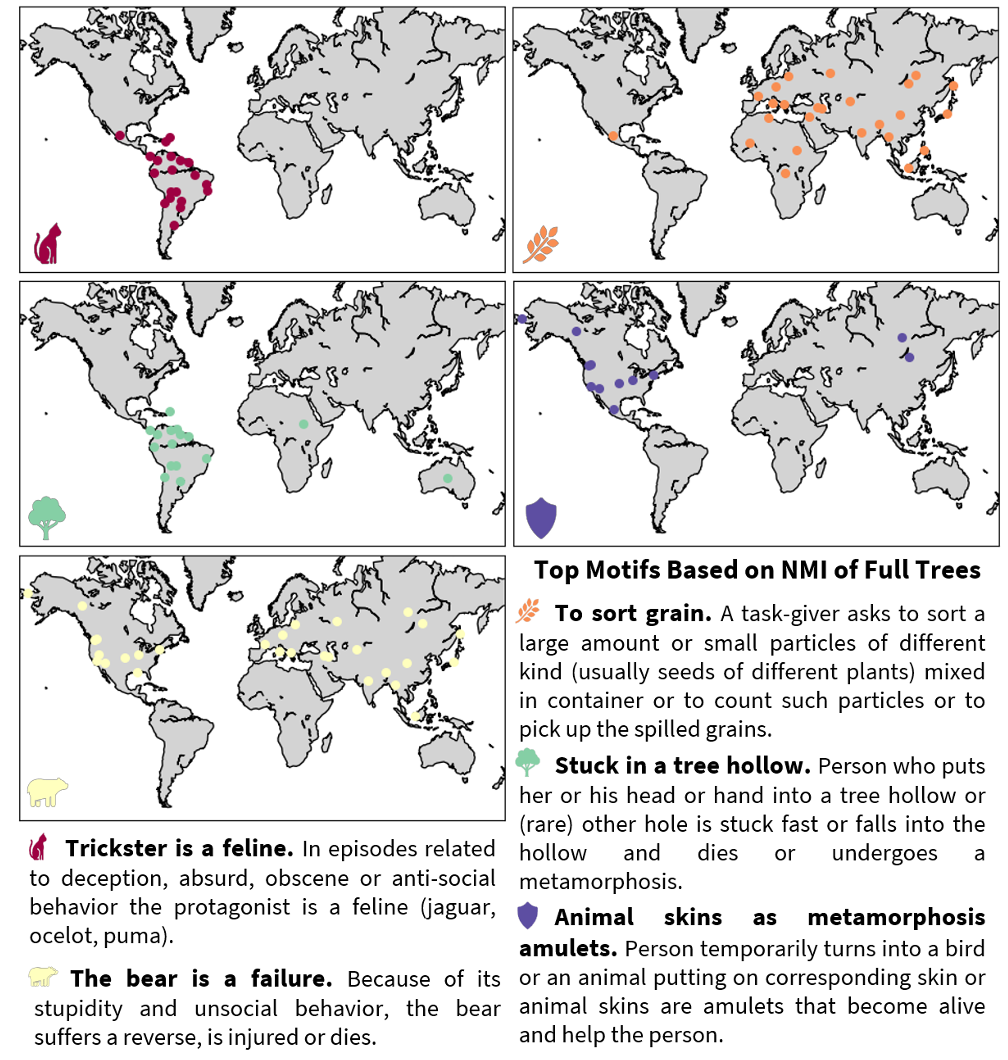
Mathematical properties of allele-sharing dissimilarities
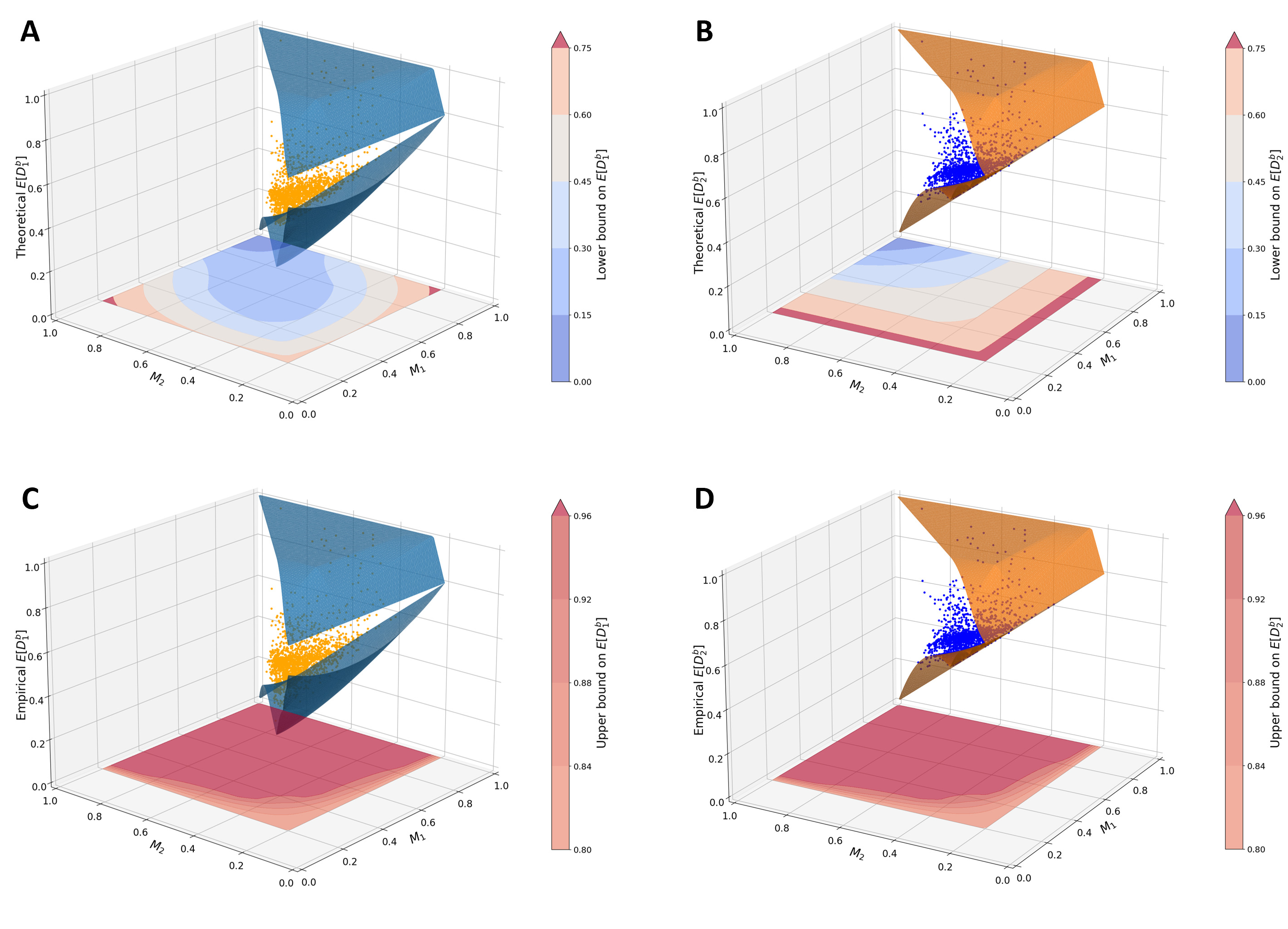
I derive and analyze the mathematical properties of variants of allele-sharing dissimilarity measures (both within and between populations) as functions of allele-frequency distributions, providing guidance on their appropriate use in empirical studies.
Alignment of inferred latent ancestries in multiple population structure analysis results
I tackle cluster misalignment in population structure inference, where the inferred latent ancestries (clusters) vary across runs and choices of number of clusters (K>). I developed Clumppling to align multiple results, consolidate distinct solutions (modes), and clarify correspondence among inferred clusters. My tool visualizes the inferred population structure across K with multipartite graphs of bar plots and explicit alignment connections, enabling consistent, interpretable comparisons.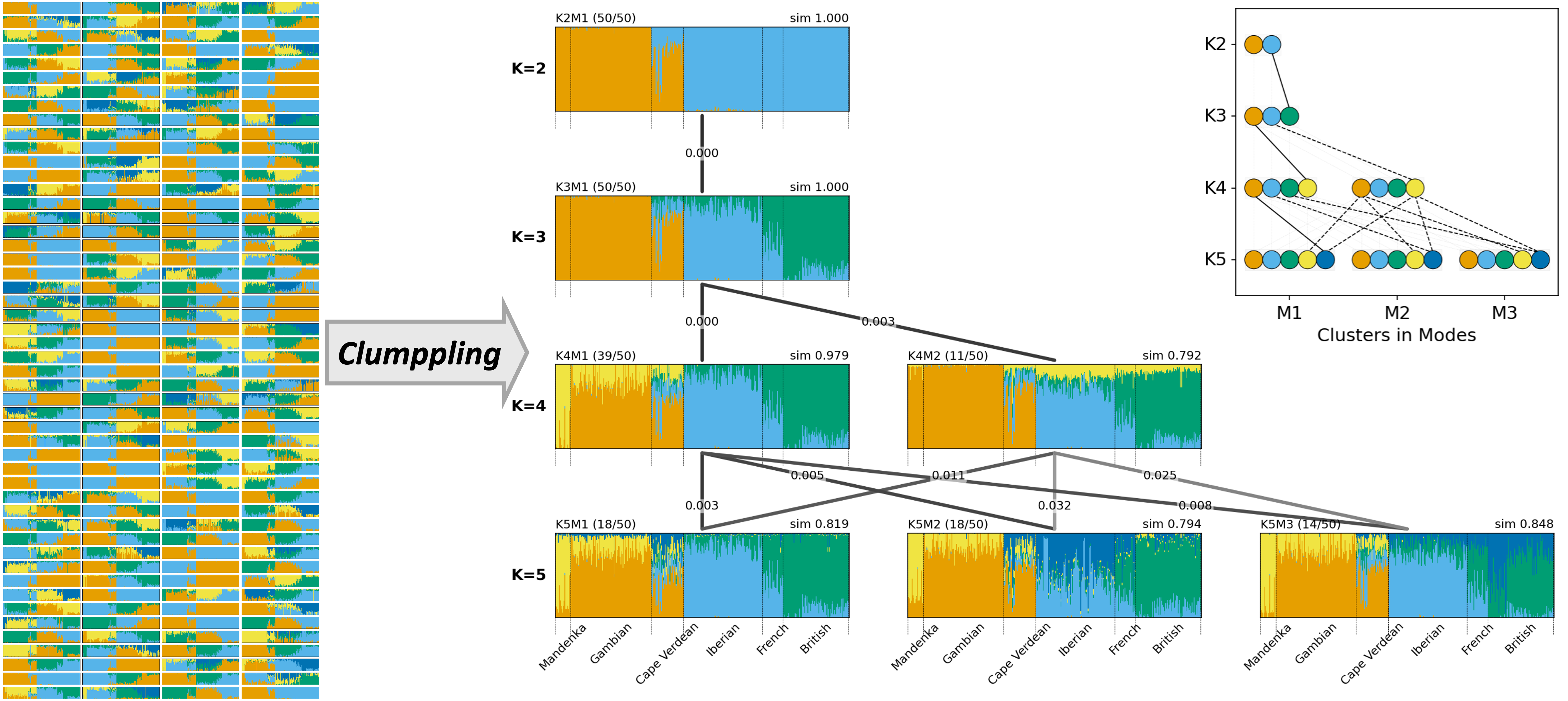
ML-based methods for multivariate genetic association analysis
I devise methods to analyze genetic association signals and better characterize the genetic architecture of complex traits. I developed ML-MAGES, which uses neural-network-based supervised learning to account for LD-induced inflation in single-trait, SNP-level genetic associations, uses an infinite mixture model to categorize multi-trait association patterns, and aggregates and visualize association signals at gene-level. This approach substantially improves computational efficiency with large LD matrices and nonlinear inflation, achieves effect-size shrinkage comparable to or better than existing methods, and enables data-driven inference of an arbitrary number of association patterns in high-dimensional multi-trait settings. 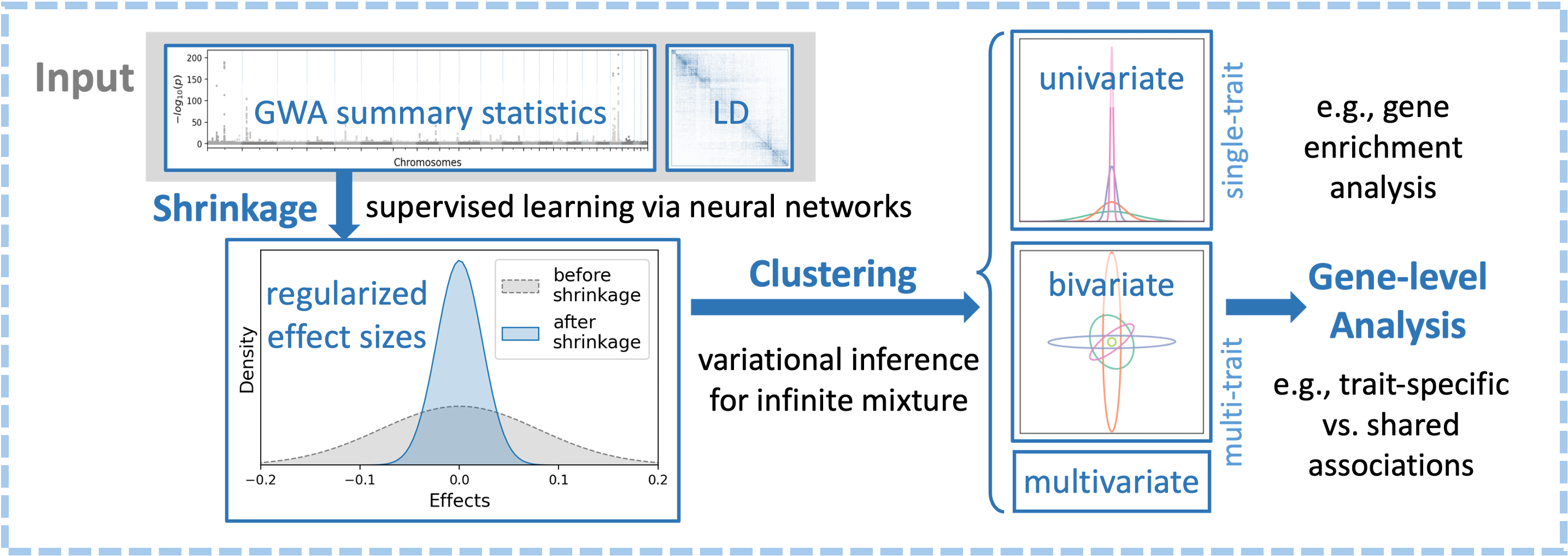
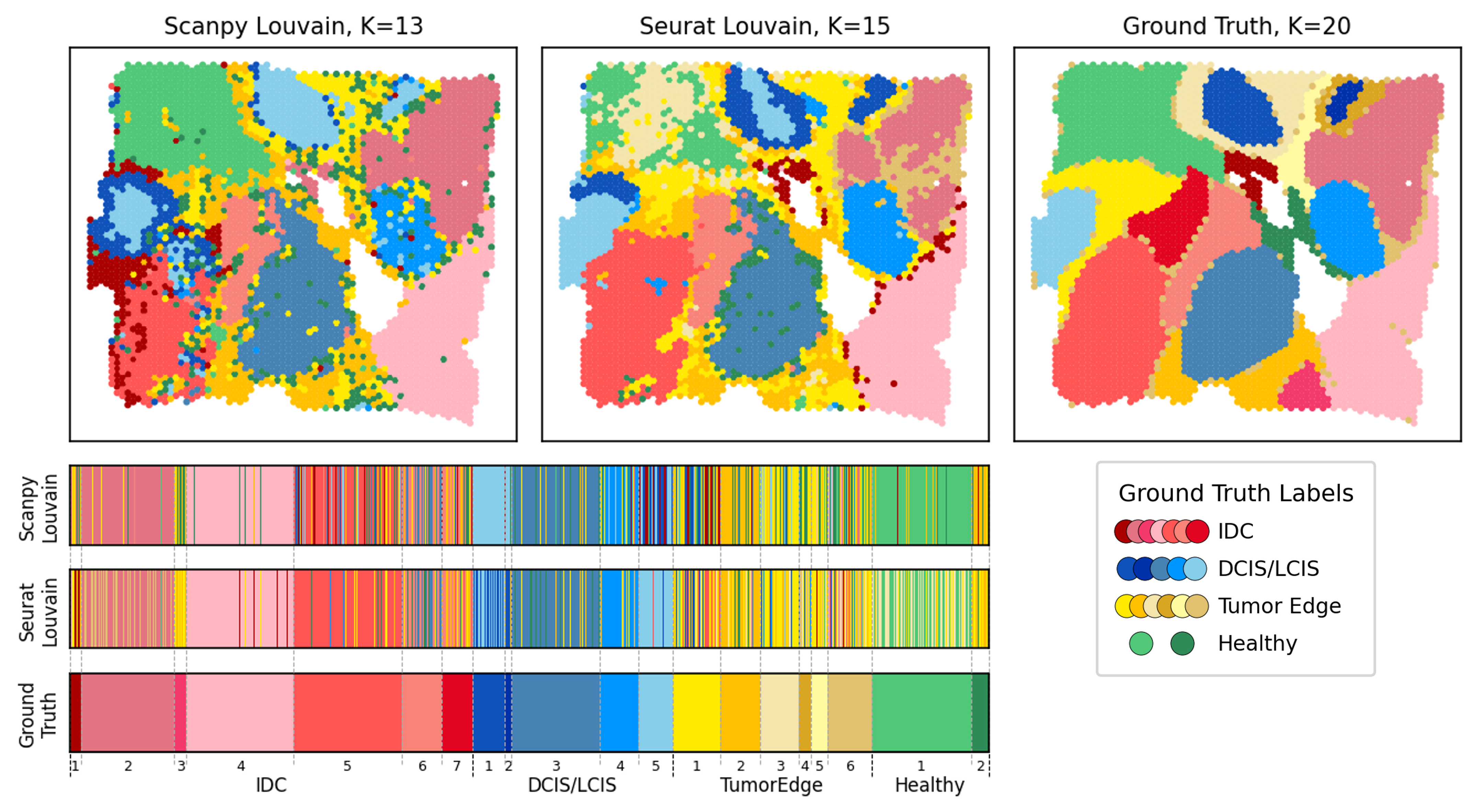
Clustering alignment to complement and advance single-cell clustering
I extend the clustering alignment framework to the single-cell analysis pipeline. Specifically, I develop new metrics for feature-level interpretation to identify cluster-informative genes, and provide measures of clustering consistency for comparing results from different clustering models. The alignment-enabled single-cell analysis pipeline streamlines robust clustering results evaluation, straightforward model comparison, and gene selection for downstream analyses.
publications
Predicting inpatient glucose levels and insulin dosing by machine learning on electronic health records
, ,medRxiv, 2020
Diffusion histology imaging combining diffusion basis spectrum imaging (DBSI) and machine learning improves detection and classification of glioblastoma pathology
, , , , , , , , , , , , , , , , , ,Clinical Cancer Research, 2020
Effects of cultural transmission of surnaming decisions on the sex ratio at birth
,Theoretical Population Biology, 2021
Extracting hierarchical features of cultural variation using network-based clustering
, ,Evolutionary Human Sciences, 2022
Deconvoluting complex correlates of COVID-19 severity with a multi-omic pandemic tracking strategy
, , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , ,Nature Communications, 2022
A Dirichlet model of alignment cost in mixed-membership unsupervised clustering
, ,Journal of Computational and Graphical Statistics, 2022
When is the allele-sharing dissimilarity between two populations exceeded by the allele-sharing dissimilarity of a population with itself?
, , ,Statistical Applications in Genetics and Molecular Biology, 2023
Clumppling: cluster matching and permutation program with integer linear programming
, ,Bioinformatics, 2024
Combinatorics of a dissimilarity measure for pairs of draws from discrete probability vectors on finite sets of objects
, ,arXiv, 2024
ML-MAGES: A Machine Learning Framework for Multivariate Genetic Association Analyses with Genes and Effect Size Shrinkage
, ,Research in Computational Molecular Biology, 2025
Using mathematical constraints to explain narrow ranges for allele-sharing dissimilarities
, ,Theoretical Population Biology, 2025
KAlignedoscope: an interactive visualization tool for aligned clustering results
, ,(under review), 2025
ML-MAGES enables multivariate genetic association analyses with genes and effect size shrinkage
, ,Genome Research, 2025
talks
A Dirichlet model of alignment cost in mixed-membership clustering results of ancestry inference.
Published:
Discussing a Dirichlet model for clustering in population structure analysis.
Clumppling: a new method for aligning replicate solutions in population structure analysis.
Published:
Presenting Clumppling, a method of clustering alignment improving population structure analysis.
ML-MAGES: A machine learning framework for multivariate genetic association analyses with genes and effect size shrinkage.
Published:
Introducing ML-MAGES, a novel ML-based framework for analyzing genetic association signals.
Clustering alignment for single cell analyses: streamlining model comparison and revealing informative genes.
Published:
Introducing clustering alignment techniques to complement and augment single-cell clustering analysis.
teaching
Ordinary Differential Equations for Engineers (CME 102)
TA for Undergraduate Course, Stanford University, Computational and Mathematical Engineering, 2020
Three offerings: Spring 2020, Winter 2021, Spring 2022.
Linear Algebra with Application to Engineering Computations (CME 200/ME 300A)
TA for Graduate Course, Stanford University, Computational and Mathematical Engineering, 2020
Two offerings: Fall 2020, Fall 2021
Advanced MATLAB for Scientific Computing (CME 292)
Instructor for Short Course, Stanford University, Computational and Mathematical Engineering, 2022
Two offerings: Winter 2022, Winter 2023
DSCoV Workshop
Instructor for Workshop, Brown University, Data Science Institute, 2025
A workshop facing the general audience, including but not limited to undergraduate and graduate students interested in data science topics.
