- Attending Biology of Genome 2026 at CSHL in May.
- Attending SMTPB 2026 at ICERM, Brown University in June.
About me
I am a Postdoctoral Researcher at Brown University, working with Sohini Ramachandran. Before that, I received my Ph.D. in Computational and Mathematical Engineering from Stanford University in 2023, where I was supervised by Noah Rosenberg.
My research focuses on method development in population genetics, computational biology, and data science. I use computational techniques such as machine learning, optimization, network analysis, and statistical inference to analyze complex genetic and genomic data.
Last updated: April, 2026
This site is still under construction 👷🔧🔨
News
Research overview
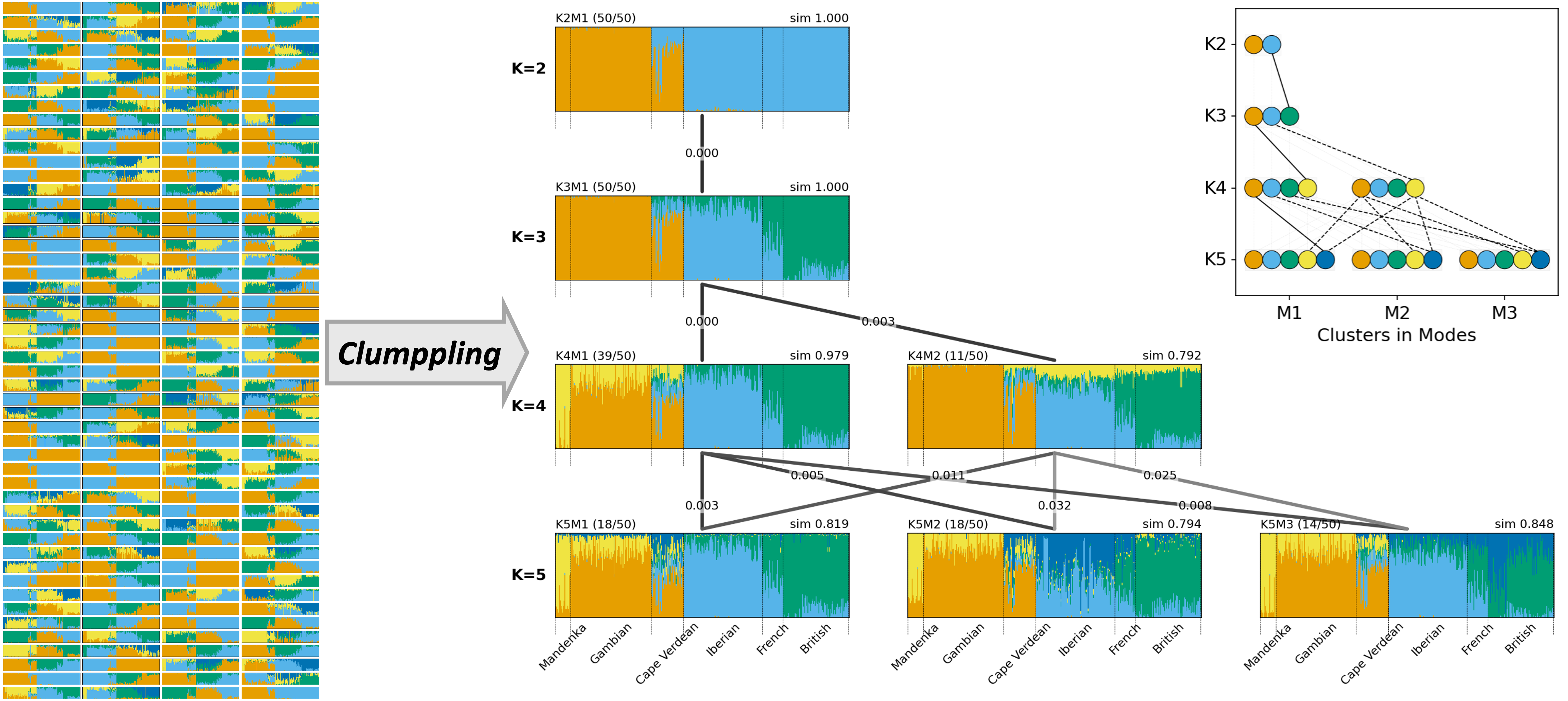
Clustering alignment for population structure analysis
Developed methods and tools for aligning latent ancestries across repeated population structure analyses so results remain interpretable across runs and choices of K.
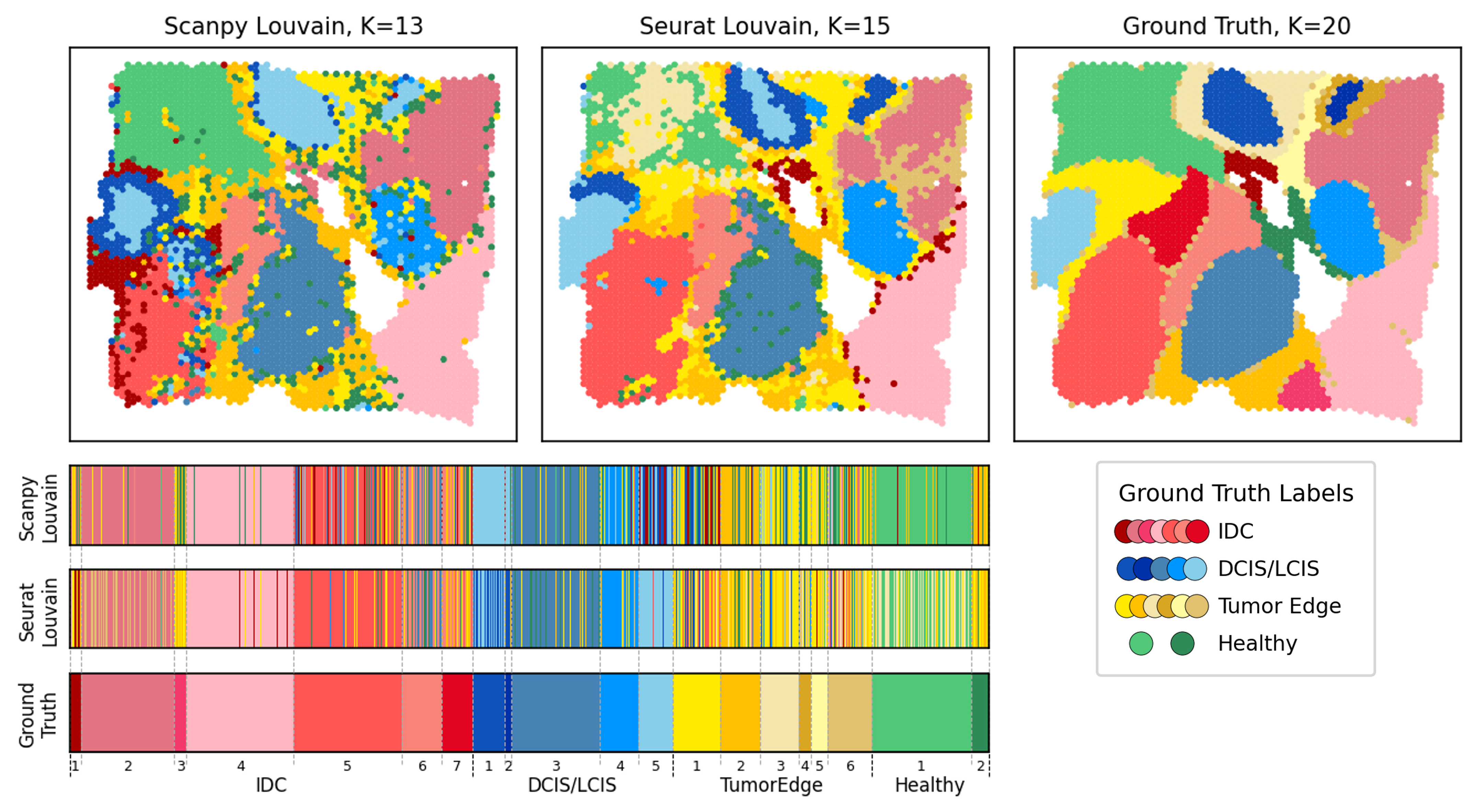
Alignment-aware single-cell clustering analysis
Extending clustering alignment ideas to the single-cell analysis pipeline to compare models, identify cluster-informative genes, and stabilize downstream interpretation.
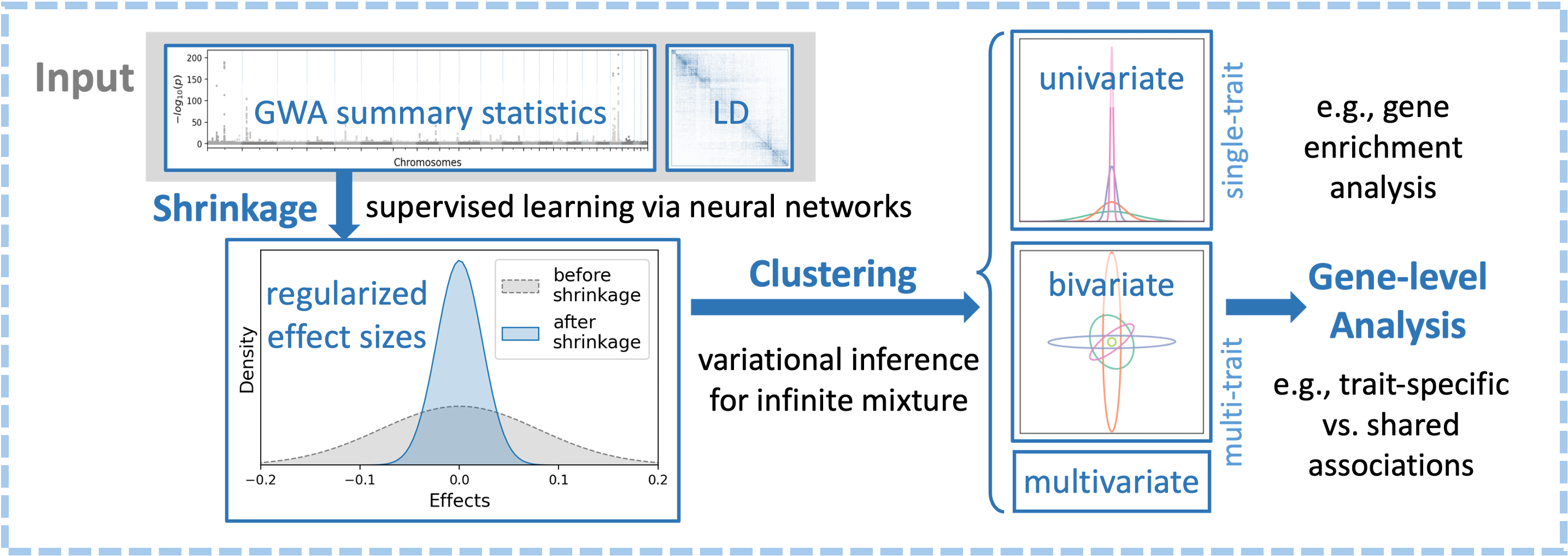
ML-based methods for multivariate genetic association analysis
Designed ML-MAGES, a machine-learning framework for shrinkage, pattern discovery, and gene-level aggregation in high-dimensional genetic association studies.
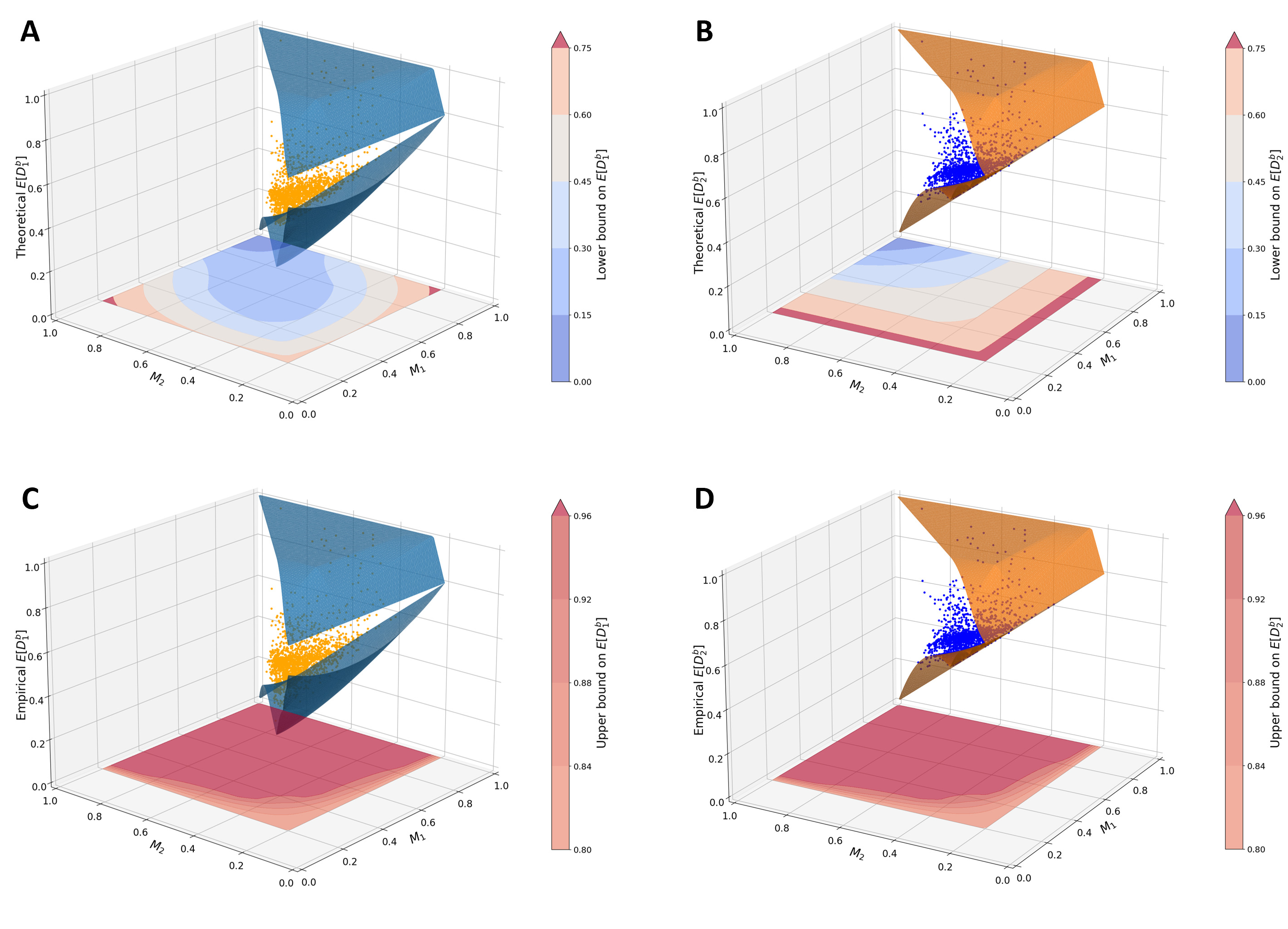
Mathematical properties of allele-sharing dissimilarities
Analyzing within- and between-population dissimilarity measures as functions of allele-frequency distributions to clarify when and how these statistics should be used.
Selected work
ML-MAGES enables multivariate genetic association analyses with genes and effect size shrinkage
Xiran Liu, Lorin Crawford, Sohini Ramachandran
Using mathematical constraints to explain narrow ranges for allele-sharing dissimilarities
Xiran Liu, Zarif Ahsan, Noah A. Rosenberg
Clumppling: cluster matching and permutation program with integer linear programming
Xiran Liu, Naama M. Kopelman, Noah A. Rosenberg