Alignment of inferred latent ancestries in multiple population structure analysis results
I tackle cluster misalignment in population structure inference, where the inferred latent ancestries (clusters) vary across runs and choices of number of clusters (K>). I developed Clumppling to align multiple results, consolidate distinct solutions (modes), and clarify correspondence among inferred clusters. My tool visualizes the inferred population structure across K with multipartite graphs of bar plots and explicit alignment connections, enabling consistent, interpretable comparisons.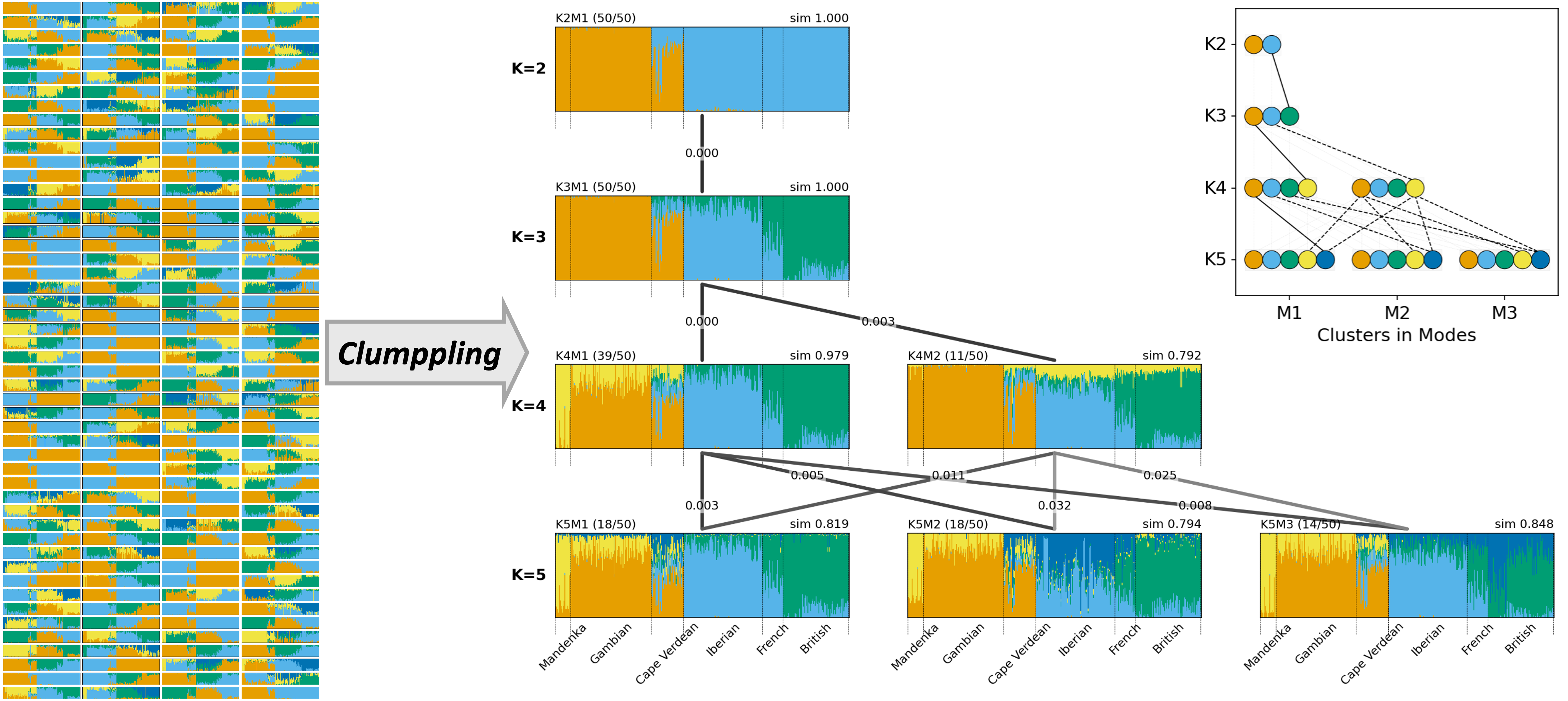
Check out Clumppling here. This is a Python-based tool to perform clustering alignment. It can be installed as a Python package via pip install clumppling.
Check out KAlignedoscope here. This is an D3.js-based visualization tool built by Avery Guo (undergrad @ Brown) for interactively exploring and displaying aligned population structure analysis results. It can be installed as a Python package via pip install kalignedoscope.
Also see here for a tutorial on the comprehensive workflow of performing population structure analysis end-to-end, from raw data preparation to creating publication-quality visualizations, using popular tools like Structure or ADMIXTURE, as well as our Clumppling and KAlignedoscope tools.
